Unveiling the Intricacies of HLA Genetic Protein Interactions
The human immune system is delicately complex, constantly modulating responses in real-time in the face of an unending, unseen assault by a myriad of pathogens. Central to this pivotal defense mechanism is the Human Leukocyte Antigen (HLA) system, a group of genes associated with a vast array of immune responses in the human body such as autoimmunity, transplantation outcomes, infectious disease susceptibilities, and drug hypersensitivity.
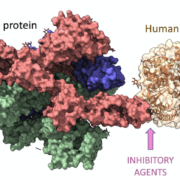
HLAs are proteins encoded by the Major Histocompatibility Complex (MHC); indeed, they are among the most polymorphic proteins encoded in the human genome. The high level of HLA polymorphism is beneficial as it increases the probability of effective immune responses to an extensive range of pathogens. However, it also presents a substantial challenge when it comes to transplantation matching, as disparities between donor and recipient HLA types may trigger transplant rejection. HLA proteins interact with a wide range of peptides derived from various pathogens, forming a complex that the immune system recognizes as ‘non-self’. These peptide-HLA complexes are then presented on the surface of cells to T cells, the vanguard of our immune system. It’s these interactions that largely dictate the course of an immune response.
To understand these intricate relationships further, scientists often use various techniques and methodologies. One crucial research method is computational modelling which provides valuable insights into the machinations at the molecular level. Scientists leverage the power of computers and algorithms to model and predict potential interactions between HLA proteins and diverse peptides. Cross-referencing these computational predictions with experimental validation bolsters the reliability of the findings. It’s remarkable to see how the nature of peptide sequence and composition influences its affinity for various HLA proteins. The binding strength and stability of the peptide-HLA complex define the efficacy of the T-cell response. Some peptides have been found to form more stable complexes with certain HLA alleles, resulting in a more robust immune response. At the same time, these binding preferences can also explain why certain HLA alleles are associated with susceptibility or resistance to particular diseases.
The HLA system’s role in drug hypersensitivity reactions also cannot be stressed enough. Some drugs interact directly or indirectly with HLA alleles, altering the usual repertoire of peptides binding to these HLAs, thereby triggering T-cell response and causing hypersensitive reactions. Predicting such interactions can assist in personalized medicine, helping avoid adverse drug reactions in individuals harboring certain HLA types. Understanding this complex web of interactions and responses within the HLA system is crucial in many areas, especially in honing therapeutic strategies against an array of diseases.
While HLA research field is riddled with puzzles ranging from peptide presentation to T-cell activation, these complexities make it an exciting and promising domain of exploration for scientists. In this era of personalized medicine, understanding the HLA system and its countless interactions could herald the revolution in healthcare we have been waiting for.

 The Sequencing Center
The Sequencing Center
 The Sequencing Center
The Sequencing Center The Sequencing Center
The Sequencing Center The Sequencing Center
The Sequencing Center
Leave a Reply
Want to join the discussion?Feel free to contribute!