The Intricate Interplay of HLA Genes, HLA Typing, and CAR-T Cell Therapy: A Deep Dive
The field of immunotherapy has witnessed significant advancements in recent years, with chimeric antigen receptor CAR-T cell therapy emerging as a revolutionary treatment modality for certain hematological malignancies. CAR-T cells are genetically engineered to express CARs that target specific tumor antigens, enabling them to selectively eliminate cancer cells. However, the success of CAR-T cell therapy can be influenced by factors such as human leukocyte antigen (HLA) genes and the patient’s HLA type. In this blog post, we will delve into the complex interplay between HLA genes, HLA typing, and CAR-T cell therapy, focusing on the importance of understanding these interactions to optimize therapeutic outcomes.
HLA Genes and HLA Typing: An Overview
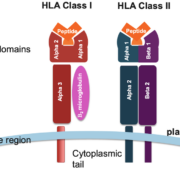
HLA genes are part of the Major Histocompatibility Complex (MHC) and encode cell surface proteins that play a crucial role in the immune system. MHC molecules present antigenic peptides to T cells, enabling the immune system to differentiate between self and non-self cells. HLA genes are classified into three groups: class I (HLA-A, HLA-B, and HLA-C), class II (HLA-DP, HLA-DQ, and HLA-DR), and class III. HLA class I and II molecules are involved in antigen presentation to CD8+ and CD4+ T cells, respectively.
Given the high degree of polymorphism in the HLA gene loci, HLA typing is essential for determining an individual’s HLA profile, which can impact the efficacy of immunotherapies such as CAR-T cell therapy. HLA typing is routinely performed for hematopoietic stem cell transplantation to identify compatible donors and minimize the risk of graft-versus-host disease (GVHD).
HLA Genes and CAR-T Cell Therapy: The Connection
The success of CAR-T cell therapy depends on the ability of CAR-T cells to recognize and eliminate cancer cells without causing off-target effects or being rejected by the host’s immune system. HLA genes play a pivotal role in this process due to their involvement in antigen presentation and T-cell recognition.
- HLA restriction and T cell receptor (TCR) specificity: CAR-T cells rely on the specificity of their TCRs to recognize tumor antigens presented by HLA molecules. As TCRs are HLA-restricted, they can only recognize antigens presented by a specific HLA molecule. This implies that the efficacy of CAR-T cell therapy can be influenced by the patient’s HLA type. For instance, a CAR-T cell therapy targeting a tumor antigen presented by HLA-A*02:01 would be less effective in patients who do not express this specific HLA allele.
- HLA loss and immune evasion: Cancer cells can evade immune surveillance by downregulating or losing HLA expression, rendering them “invisible” to T cells. This immune evasion mechanism can limit the effectiveness of CAR-T cell therapy, as CAR-T cells are unable to recognize and eliminate cancer cells that lack HLA expression.
- Alloreactivity and graft-versus-leukemia (GVL) effect: In the context of allogeneic CAR-T cell therapy, donor-derived CAR-T cells can recognize recipient HLA molecules as foreign and mount an alloreactive response. This phenomenon, known as the GVL effect, can contribute to the elimination of residual cancer cells. However, it can also result in GVHD, a potentially life-threatening complication. HLA typing is crucial for identifying compatible donors and mitigating the risk of GVHD.
Optimizing CAR-T Cell Therapy through HLA Typing
Understanding the complex interplay between HLA genes and CAR-T cell therapy can help optimize treatment strategies and enhance therapeutic outcomes. Here are some approaches that leverage HLA typing to improve CAR-T cell therapy:
- Personalized CAR-T cell therapy: Developing personalized CAR-T cell therapy based on a patient’s HLA type can enhance the specificity and efficacy of treatment. By designing CARs that target tumor antigens presented by the patient’s specific HLA molecules, CAR-T cells can effectively recognize and eliminate cancer cells, reducing the likelihood of immune evasion.
- HLA-edited CAR-T cells: Gene editing technologies, such as CRISPR/Cas9, can be employed to modify CAR-T cells to express specific HLA molecules compatible with the patient’s HLA type. This approach can reduce the risk of alloreactivity and GVHD in the context of allogeneic CAR-T cell therapy. Moreover, HLA editing can also prevent the rejection of CAR-T cells by the host’s immune system, enhancing their persistence and therapeutic efficacy.
- Targeting HLA loss in cancer cells: Strategies that target the mechanisms underlying HLA loss in cancer cells can help restore HLA expression and enhance the effectiveness of CAR-T cell therapy. For example, small molecules that inhibit the activity of histone deacetylases (HDACs) can upregulate HLA expression in cancer cells, rendering them susceptible to CAR-T cell-mediated killing.
- TCR-mimic CARs: An alternative approach to enhancing the efficacy of CAR-T cell therapy is the development of TCR-mimic CARs that recognize peptide-HLA complexes rather than native tumor antigens. TCR-mimic CARs can bypass the limitations associated with HLA restriction and immune evasion, enabling CAR-T cells to target a broader range of cancer cells.
Conclusion
The intricate interplay between HLA genes, HLA typing, and CAR-T cell therapy highlights the importance of understanding these interactions to optimize therapeutic outcomes. By leveraging HLA typing and incorporating novel strategies, such as personalized CAR-T cell therapy, HLA-edited CAR-T cells, targeting HLA loss, and TCR-mimic CARs, researchers and clinicians can enhance the efficacy and safety of CAR-T cell therapy for patients with hematological malignancies and potentially other types of cancer. As our understanding of these complex interactions continues to evolve, so too will the potential for more precise and effective immunotherapies.



 The Sequencing Center
The Sequencing Center
Leave a Reply
Want to join the discussion?Feel free to contribute!